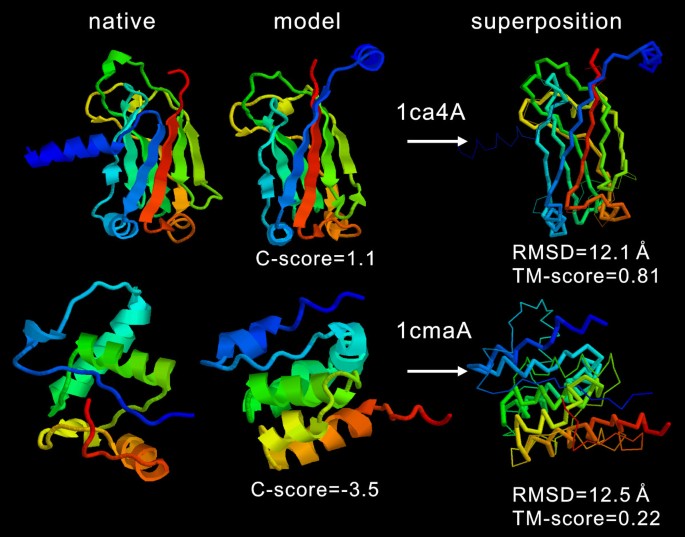
Examples of I-TASSER models from three independent benchmark sets. The... | Download Scientific Diagram
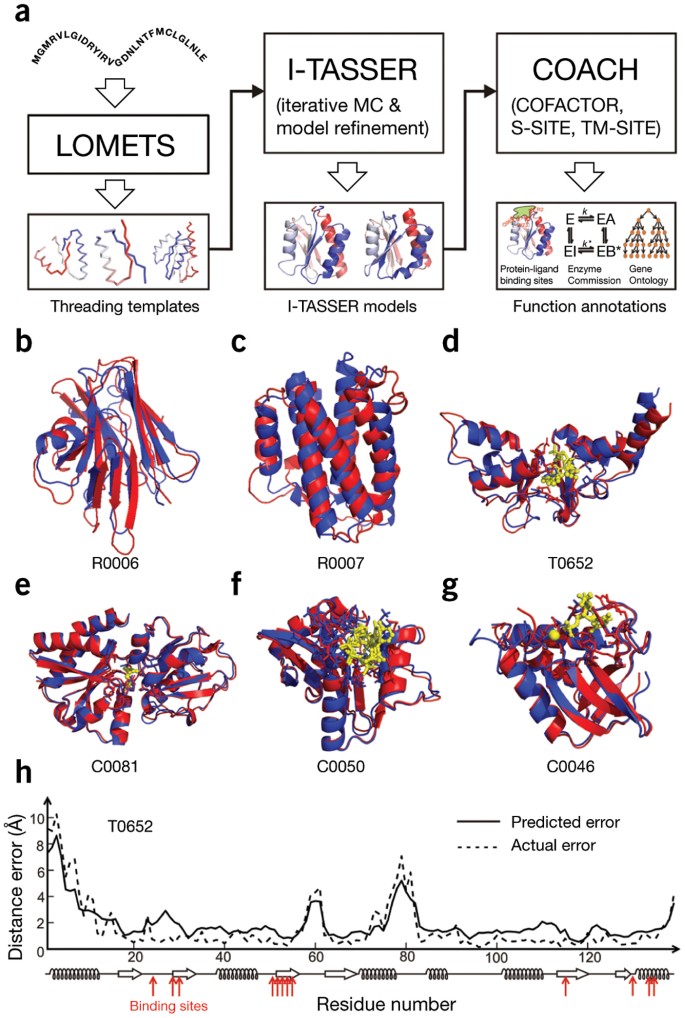
![PDF] Folding non-homologous proteins by coupling deep-learning contact maps with I-TASSER assembly simulations | Semantic Scholar PDF] Folding non-homologous proteins by coupling deep-learning contact maps with I-TASSER assembly simulations | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/57438546e23780b3b7b838e562b4817a27dc6bb2/4-Figure2-1.png)
PDF] Folding non-homologous proteins by coupling deep-learning contact maps with I-TASSER assembly simulations | Semantic Scholar
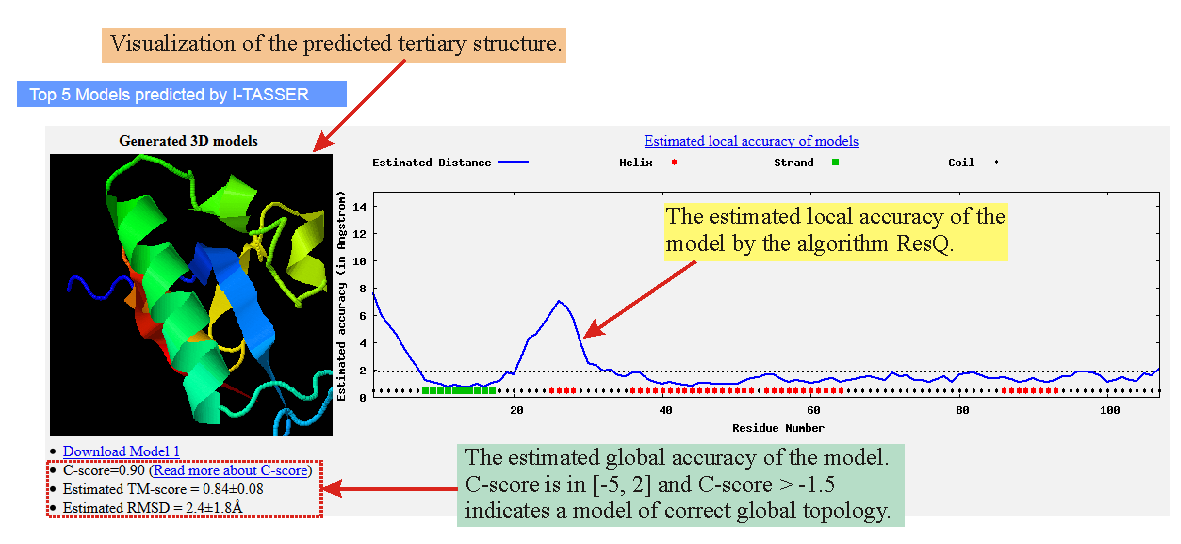
![PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/682fc6a2dcf6edaadacb1a2f90fdd137aef28d15/2-Figure1-1.png)
PDF] I-TASSER server: new development for protein structure and function predictions | Semantic Scholar

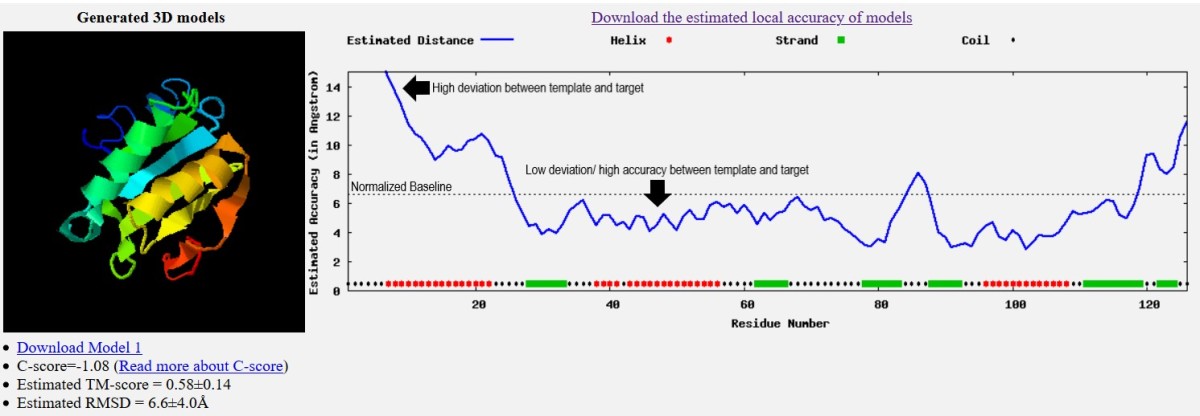
I-TASSER | Protein 3D structure prediction | Results Analysis | Lecture 6 Part 3| Dr.Muhammad Naveed - YouTube

Three-dimensional structure of proteins obtained from I-TASSER a ESAT-6... | Download Scientific Diagram
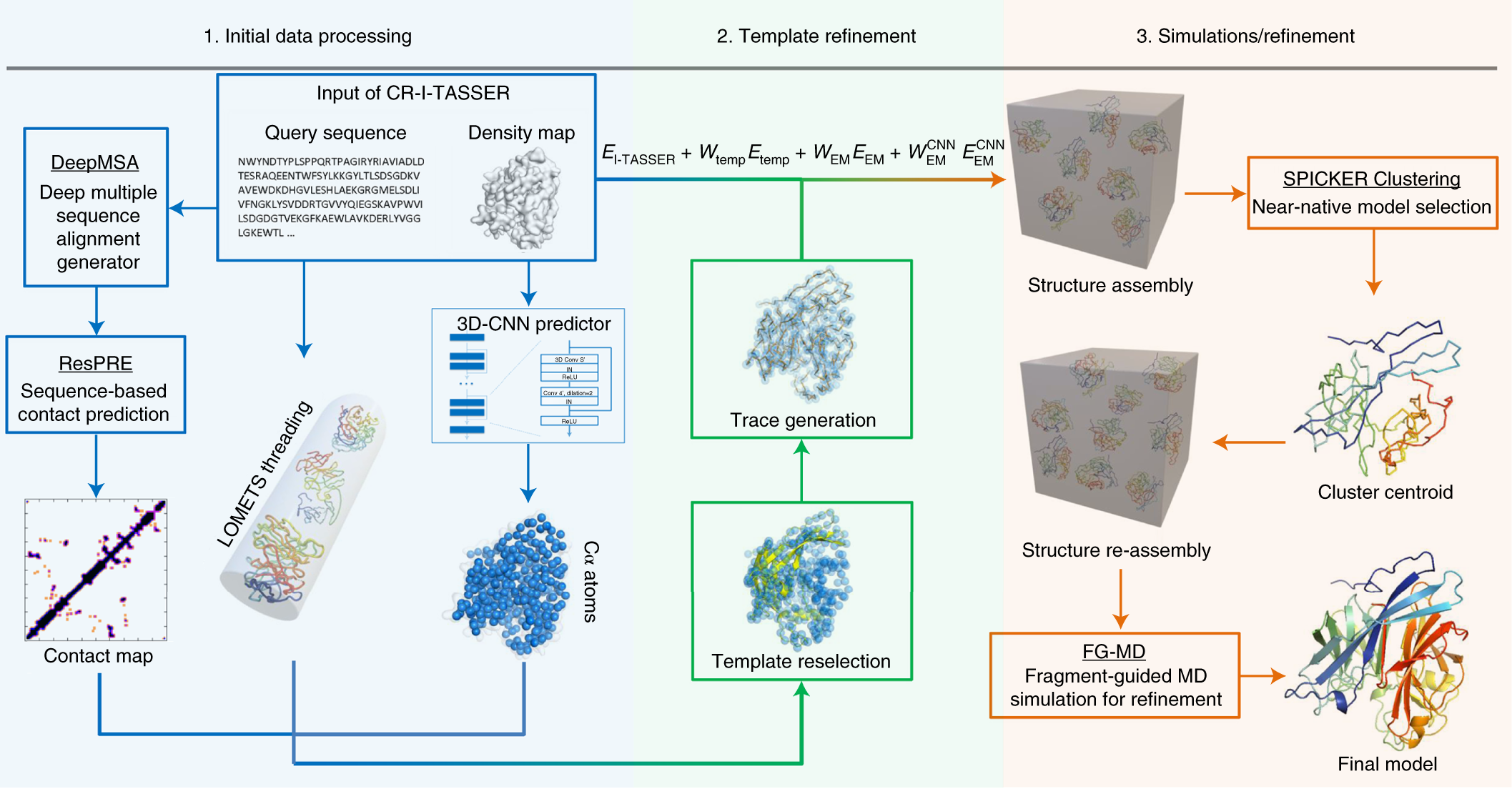
CR-I-TASSER: assemble protein structures from cryo-EM density maps using deep convolutional neural networks | Nature Methods

Computational Protein Design and Large-Scale Assessment by I-TASSER Structure Assembly Simulations - ScienceDirect

Template‐based modeling and free modeling by I‐TASSER in CASP7 - Zhang - 2007 - Proteins: Structure, Function, and Bioinformatics - Wiley Online Library

I-TASSER-MTD: a deep-learning-based platform for multi-domain protein structure and function prediction | Nature Protocols

Folding non-homologous proteins by coupling deep-learning contact maps with I-TASSER assembly simulations - ScienceDirect