Quantitative label-free mass spectrometry using contralateral and adjacent breast tissues reveal differentially expressed proteins and their predicted impacts on pathways and cellular functions in breast cancer - ScienceDirect

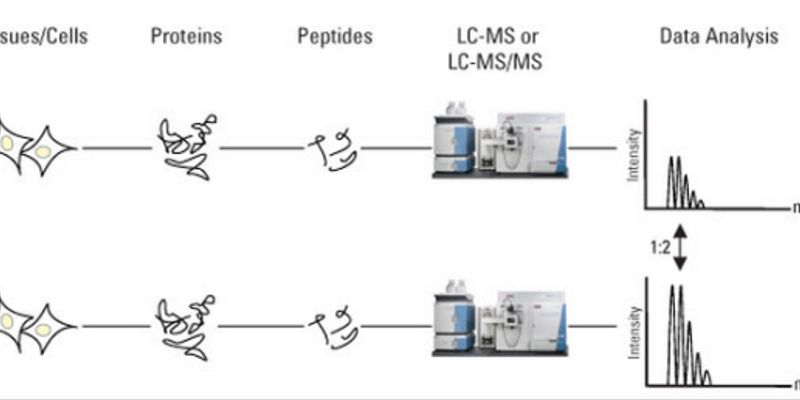
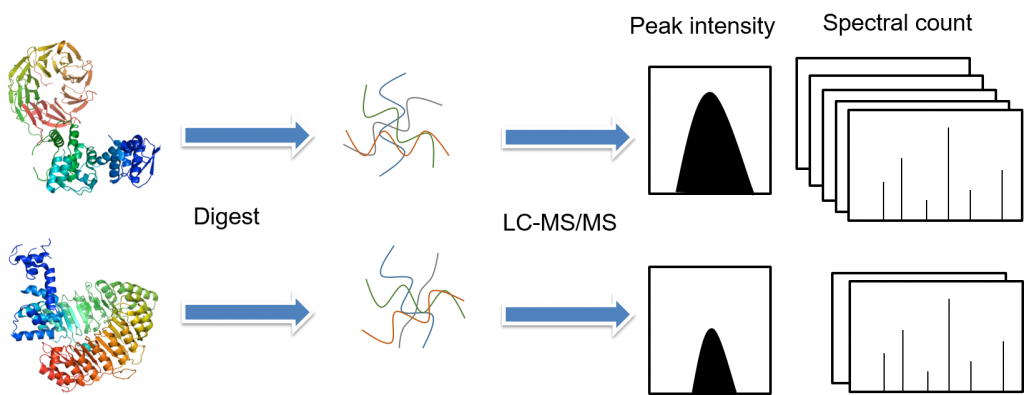
Differences between label free quantification (A) and quantification by... | Download Scientific Diagram

Robust Summarization and Inference in Proteome-wide Label-free Quantification - Molecular & Cellular Proteomics

Comparison of label-free and label-based strategies for proteome analysis of hepatoma cell lines - ScienceDirect

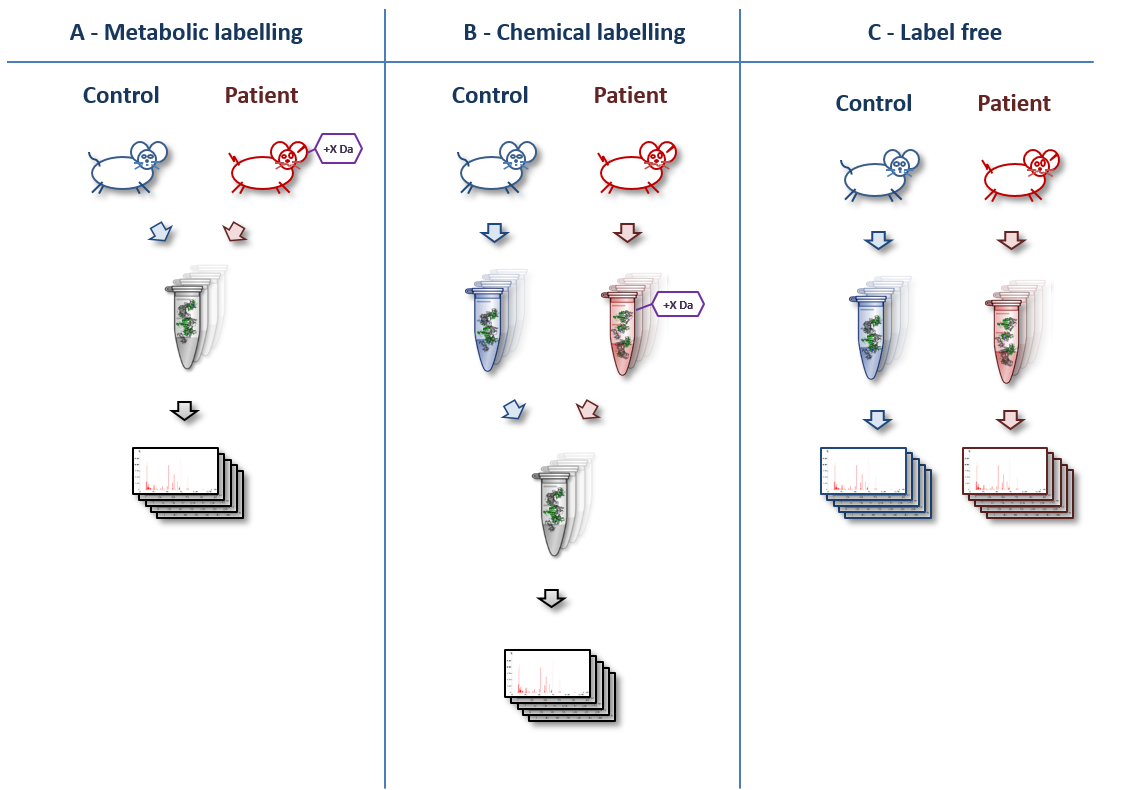
Mass spectrometry–based measurements. (a) Sample processing. Label-free... | Download Scientific Diagram

Label-free quantitative proteomics of the lysine acetylome in mitochondria identifies substrates of SIRT3 in metabolic pathways | PNAS

Accurate Proteome-wide Label-free Quantification by Delayed Normalization and Maximal Peptide Ratio Extraction, Termed MaxLFQ* - Molecular & Cellular Proteomics

IonStar enables high-precision, low-missing-data proteomics quantification in large biological cohorts | PNAS